|
I wouldn't assume they change drastically, but it's not the identical values either. You can figure out an estimate value by virtually titrating the tripeptide and figuring out after what step an isolectric point likely situates: Where a,b,c are the number of tyrosine, trytophan and cystine residues per mole of protein and E residue are the molar extinction rated of the residue at the wavelength used (280 nm). About Relative Standard Deviation Calculator Basic AA: pI (pKa R group + pKa Amino. All rights reserved.I think there is not enough data to provide an exact answer theoretically. The calculations is as follows: EM,Gdn-HClaEM,Tyr + bEM,Trp + cEM,Cys. 2: Peptide bond forms between amino acid an antigen is a protein. To assist in determining similarity we define two classes of acids. If additional acidic or basic groups are present as side-chain functions, the pI is the average of the pK a s of the two most similar acids. 
You can also learn more detailed characters such as isoelectric point. Thus, the pI for alanine is calculated to be: (2.34 + 9.69)/2 6.02, the experimentally determined value. Use the scrambler tool to randomly shuffle peptide or protein sequences.Ĭopyright © 2012-2023 . Use our Peptide Molecular Weight Calculator to check the molecular weight of your peptide. Try the sequence splitter tool for fragmenting sequences of proteins and peptides with optional overlap. Peptide with Phosphoserine: Arg-Arg-Ala-Ser(PO3)-Pro-Val-Ala Also on this site Peptide with isotopically labeled Valine and Isoleucine: Peptide with disulfide bridge (1-6), capped at C-terminal (amide): Cyclic CYFQNCPRG-NH2 Peptide with Biotin on N-terminal: Biotin-YGRKKRRQRRR 
Peptide with two disulfide bridges: ACDCRGDCFCG Head-to-tail cyclic peptide: cyclo(GRGDSP) The presence of additional acidic groups (such as the side chains of aspartic and glutamic acid), which have low pKa values, will lower a proteins pI. Linear peptide in three-letter code: Arg-Gly-Asp-Cys it is not a trivial task to determine the primary structure of such compounds. Linear peptide in one-letter code: KYICNSSCM The conformational flexibility of peptide chains is limited chiefly to. Please note: this is a rapidly changing project in development look, feel, and functionality may and will change. Take a look at the following posts for demonstration of what the calculator is capable of:įor a quick sample of the output screen, you can load a random peptide from the list of peptides the analytical tool has calculated before. At pH 3.52, the H+ concentration is high (low pH more acidic more H+). How do you find the pi value of cysteine For cysteine, pI 5.02. This is an equation of the form y mx where the slope m is pi (). The formula you graphed is circumference pi diameter. The peptide molecular weight calculator will display average and monoisotopic mass of the molecule, as well as a table of mass divided by charge values, both in positive and negative scan modes, which is useful for mass spec analysis. Peptide Calculator Peptide Molecular Weight Calculator Sequence How to use this peptide analytical tool Simply type in, or copy and paste, peptide or protein fragment amino-acid sequence, including modifications, spacers, or special termini, and press the Calculate button. This ratio, which is the slope of the line, is called pi (). Simply type in, or copy and paste, peptide or protein fragment amino-acid sequence, including modifications, spacers, or special termini, and press the “Calculate” button. Since the p I is the p H at which the amino acid has no overall net charge, you need to average the p K a values relevant to the protonation/deprotonation of the form with no net charge. As Dawid says, if youre going to use Ni-NTA purification and your protein has a pI 6, then. acid sequence of analyzed protein (peptide) or multiple sequences in fasta format. The choice of going up or down on the pH scale depends on what you want to do with your protein. 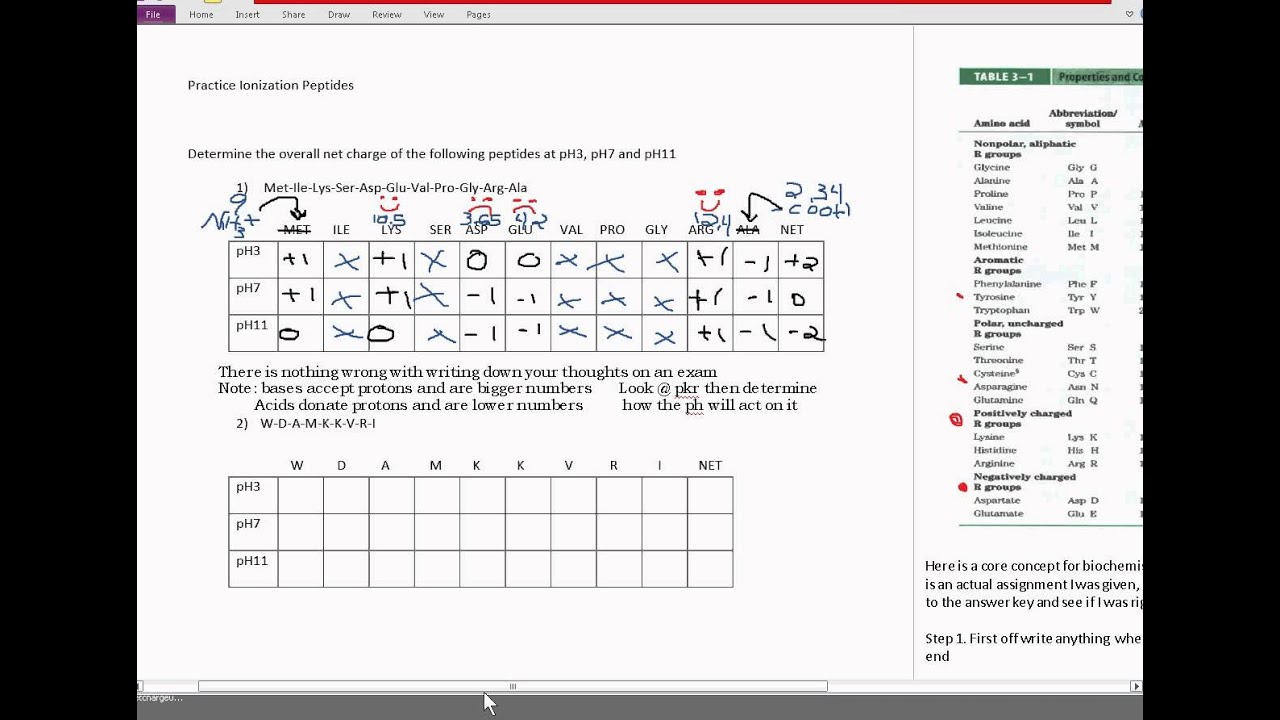
pKa 1 -carboxyl group, pK a 2 -ammonium ion, and pK a 3 side chain group. Online calculation (prediction) of theoretical isoelectric point (pI. Sequence How to use this peptide analytical tool The pK a values and the isoelectronic point, pI, are given below for the 20 -amino acids. Transcribed image text: The structure of peptide is given by plaz 10.5 Ho O H H ha Pk 9.6 pka2.0 HOT Pe: 10.5 are they going Pka 6.0 peptide going to take a proton from the solution or going to donate a proton to the solution which can i figur out with the value from pka.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
